|
|
| Fast-reaction-transport modeling - MATLAB code |
This MATLAB code is provided as supplementary material to the following article:
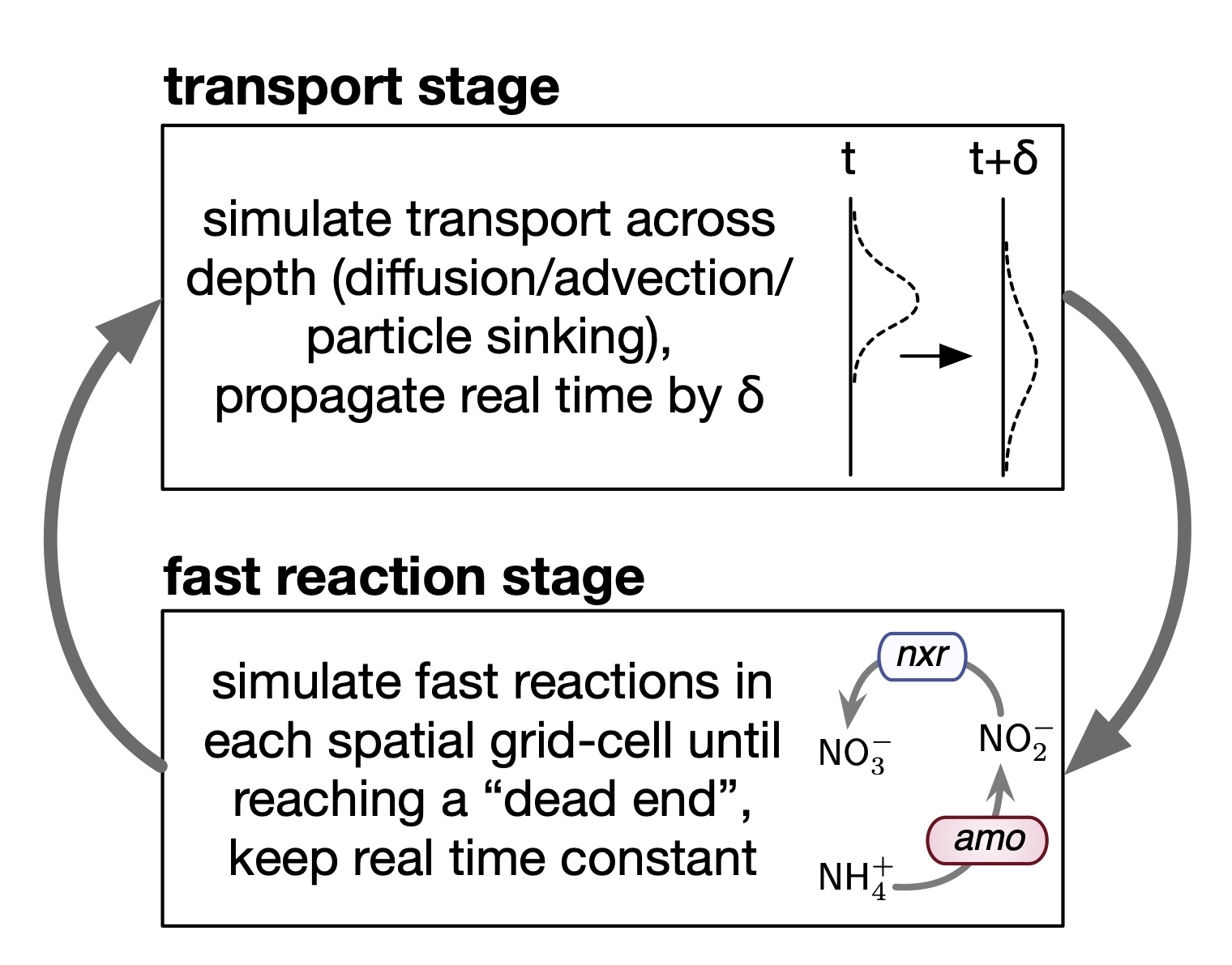
Fast-reaction-transport (FRT) biogeochemical models constitute a special limit of reaction-transport partial differential equation models, where microbial population dynamics and reaction kinetics operate at much faster time scales than the physical transport/mixing processes.
They have the benefit of not requiring any knowledge of microbial population dynamics, physiology, species composition or kinetics, thus greatly reducing the number of free parameters needed for making biogeochemical predictions.
Multiple independent code bases are provided here, for each of the following tasks:
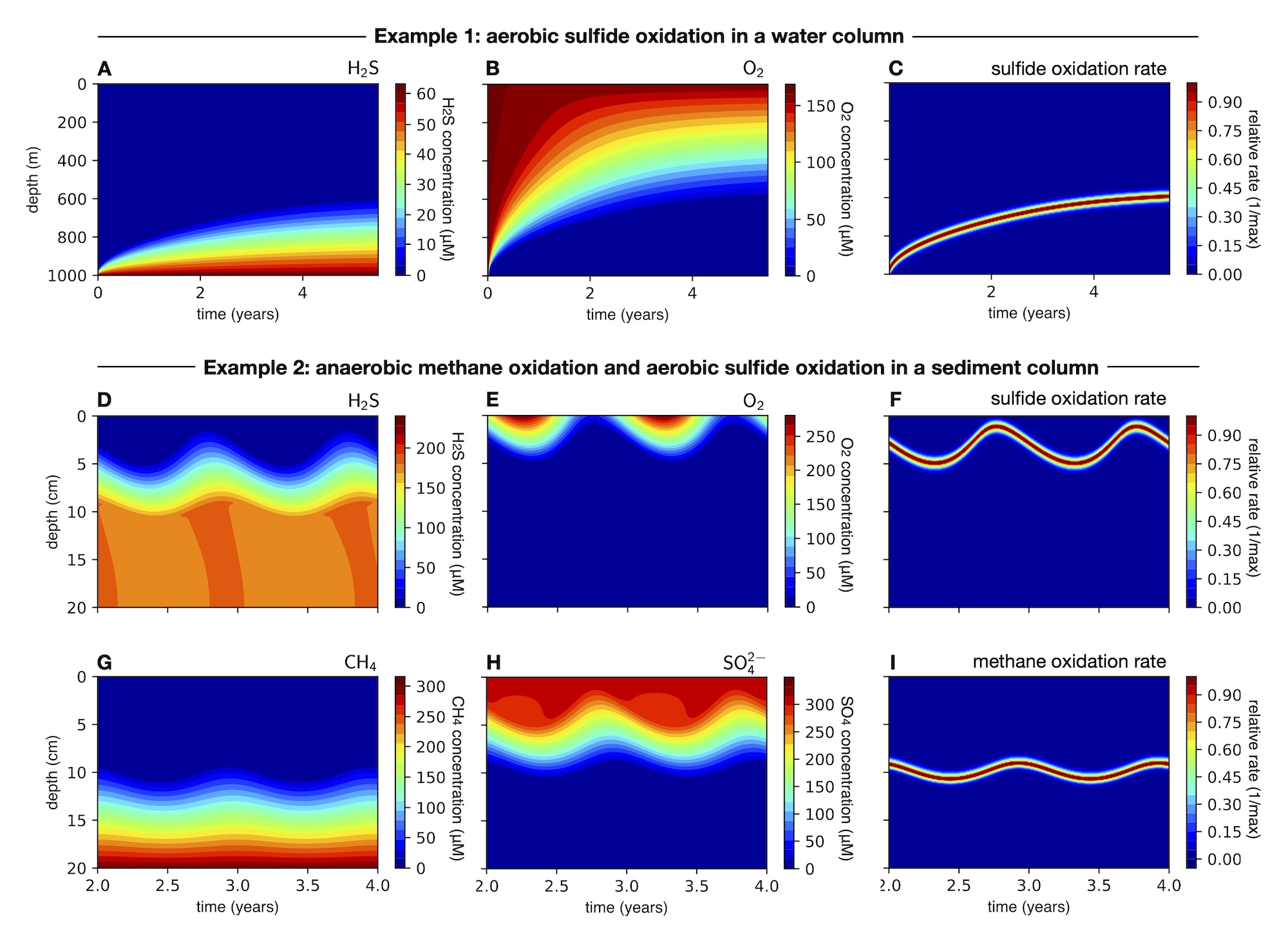
- A simple FRT model demonstrating sulfide and oxygen dynamics in a poorly mixed water column, where sulfide diffusing upwards from the bottom is oxidized aerobically using oxygen diffusing downwards from the top.
- A simple FRT model demonstrating methane, sulfide, sulfate and oxygen dynamics in a sediment column, where methane is oxidized anaerobically using sulfate, thus producing sulfide, and the produced sulfide is subsequently oxidized using oxygen at a shallower depth.
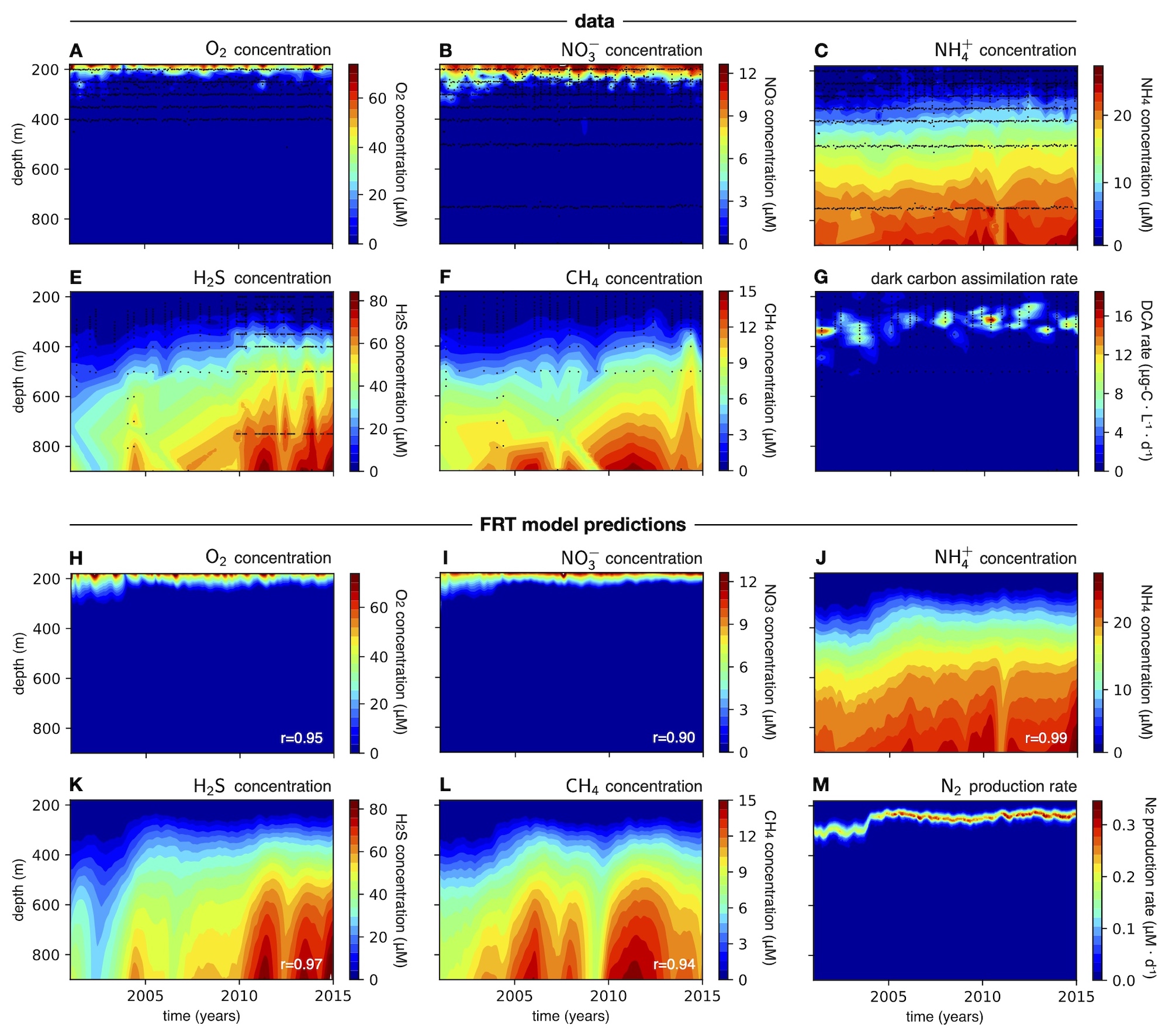
- A realistic FRT model applied to the Cariaco Basin dysphotic zone and compared to chemical time series data spanning 14 years. The chemical concentrations predicted by the model across depth and time are in exceptionally good agreement with the data, suggesting that the considered microbial metabolic rates in this system are largely diffusion-limited.
In all cases, simulations of the FRT model are performed by a custom function implemented in the MATLAB file runFRTmodel.m. This functin can handle an arbitrary number of metabolites and reactions, separate diffusion coefficients and advection speeds for each metabolite (each of which can vary with depth), and various combinations of boundary conditions (Dirichlet and/or Robin).
A regular MATLAB installation should be sufficient to run this code.
The code has been tested in MATLAB R2022a, on Mac OS 10.15.7.
Other MATLAB versions may require some slight modifications of the code.
Please consult the README files included in the downloads below for technical details and license agreements.
Downloads:
 | MATLAB code for the sulfide oxidation FRT model in a water column. |
 | MATLAB code for the methane + sulfide oxidation FRT model in a sediment column. |
 | MATLAB code and input data for FRT modeling of the Cariaco Basin water column. |
|
|
 ×
 ×
 ×
|
|
|
Louca lab. Department of Biology, University of Oregon, Eugene, USA
© 2025 Stilianos Louca all rights reserved
|
|
|



