Further Tutorials
Relative gradings and spinc structures
bfh_python computes the decomposition of HF-hat into spinc-structures and the relative Maslov grading in each spinc-structure. It also computes the relative, rational Maslov grading between torsion spinc-structures, and the ambiguity of the Maslov grading in non-torsion spinc-structures (the divisibility of the first Chern class).
A lens space
To get the relative gradings:
The first entry is the relative Maslov grading, the second is the ambiguity, and the third is the relative spinc-grading. Since the generators represent different spinc structures, the first entry is essentially meaningless here. The second is always zero, because all generators represent torsion spinc-structures. The third is the interesting one: we get five different numbers, so the generators represent five different spinc-structures.
We can get the relative rational Maslov gradings instead:
We’ve lost the decomposition into spinc-structures, but now the first entry gives us the relative rational grading between the first generator and the given one. I get 0/1, 4/5, 0/1, 2/5, and 4/5, which is correct for the relative rational gradings on L(5,2). In particular, the fourth generator represents the unique spin structure.
Σ(2,3,7)
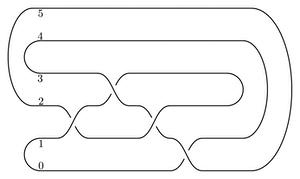
Here is the branched double cover of the (2,-3,-7) pretzel knot:
We get the relative gradings of generators as above:
There are two generators in the same grading and one in grading 1 smaller, which is correct for Σ(2,3,7).
A non-torsion spinc-structure
Here is the branched double cover of L10n59, a two-component, determinant zero link.
The output is: (0,2,[0,0]), (1,2,[0,0]), (3/2,2,[0,1]), (7/2,0,[-1,-1]), (9/2,2,[0,1]), (5/2,0,[-1,-1])
Looking at the second components, HF-hat is supported in spinc-structures with first Chern classes -2, 0, and 2. For computing the gradings, we’ve arbitrarily chosen a base generator with first Chern class ±2. The third component tells us that we have three different spinc-structures represented. In each there are two generators (sensible). In each of them, the Maslov grading difference of those generators is one: in the spinc-structure labeled [0,1] it looks like the Maslov grading difference is 9/2-3/2=3, but the Maslov grading is only well-defined mod 2.
Involutive Heegaard Floer homology
To compute HFI-hat of a manifold Y, you need to split Y=Y1∪Y2, compute P=CFD(Y1) and Q=CFD(Y2), and then invoke involutiveCx(P,Q). This can take a long time to run; you can add verbose=True as an argument to have it print its progress along the way. What the function involutiveCx is doing is documented in comments inside the function.
For example, here’s the HFI-hat of the lens space L(5,2):
(You should get 6: the conjugation-invariant spin structure contributes 2 and the other spinc-structures contribute 1 each.)
To compute HFI-hat of branched covers, there are convenience functions invOfDCov and invOfDCovSplit. The input to invOfDCov has the same format as the input to readBridgePresentation in braid.py; the docstring to readBridgePresentation is fairly thorough. So, the following computes HFI-hat of the branched double cover of the figure-8 knot:
The function invOfDCovSplit takes an integer in addition to a bridge presentation: where to split the bridge presentation when computing HFI-hat (i.e., after which crossing, from 0 to n=the number of crossings). (Sometimes, splitting in one place is much faster than splitting in another.)
Funding to support writing this documentation was provided by NSF Grant DMS-1810893. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
© 2019 - 2026 Robert Lipshitz lipshitz@uoregon.edu