research
Research in the Phillips lab spans a wide range of biological questions, from molecular genetics to evolutionary ecology. The central thread of our work is the use of genetic, experimental and theoretical approaches to address fundamental questions in evolutionary biology. We primarily use the model nematode Caenorhabditis elegans and its relatives (particularly C. remanei) for our empirical work.
Currently Funded Projects
MIRA: Systems genomics of complex traits
NIH R35GM131838
Genome engineering for a novel post-reproductive genetic screen for increased longevity
NIH R21AG086869
Caenorhabditis Intervention Testing Program at the University of Oregon
NIH U01AG045829
Caenorhabditis Intervention Testing Program Data Center
NIH U24AG056052
Research Projects
The current core research areas in the lab are:
Evolutionary genetics of aging and stress response
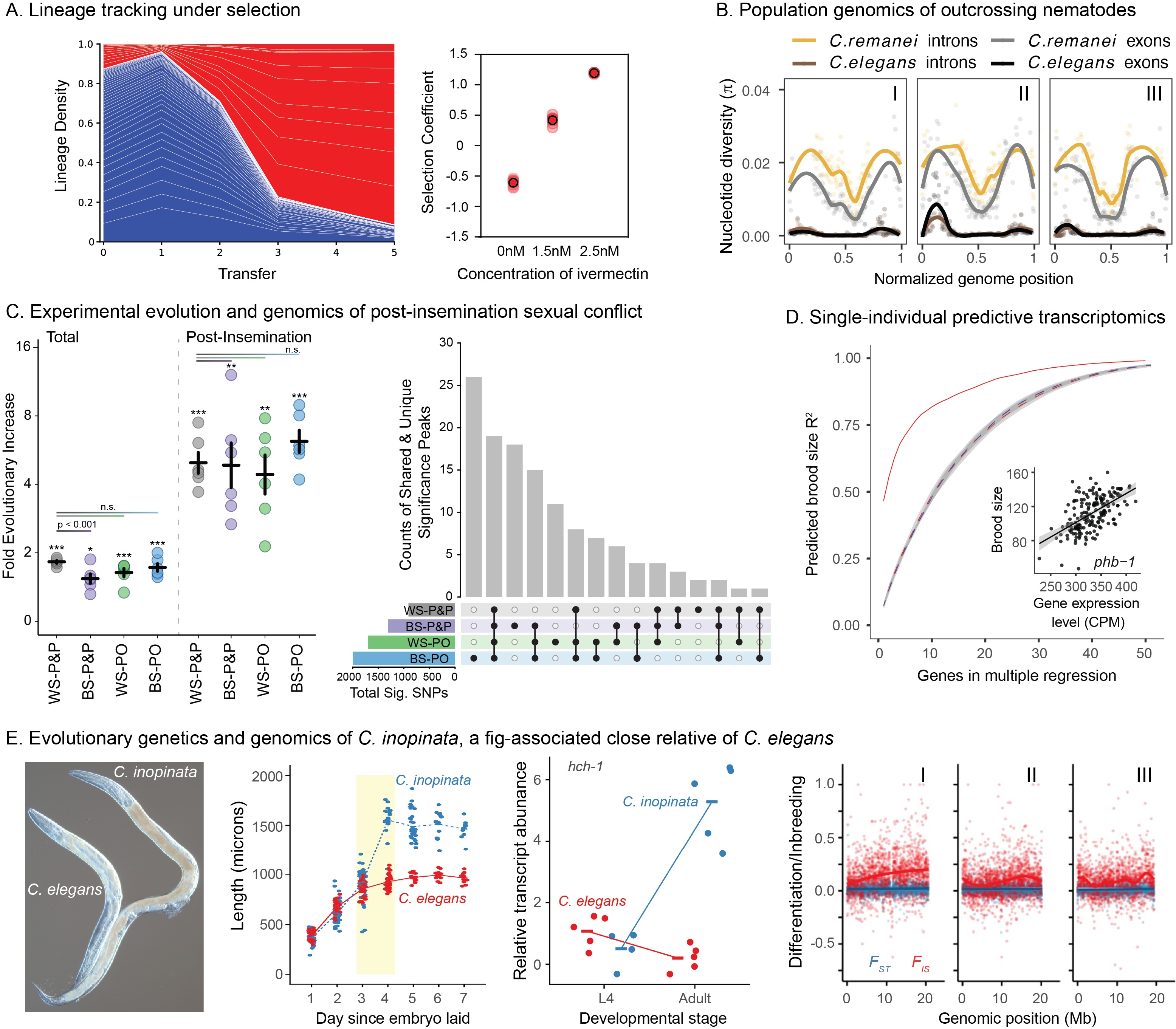
Many of the key insights into the genetic basis of aging have been made in C. elegans. We are capitalizing on this knowledge to understand the genetics of natural variation in aging and the response to a wide variety of environmental stressors (e.g., heat, oxidation, pathogens) in C. remanei. We are conducting large scale experiments selecting on the response to both chronic and acute stress. We are analyzing genetic variation both at the level of the physiological response and in the underlying signal transduction and response networks.
Compound/drug interventions that increase lifespan
Researchers have begun to identify chemical compounds that can extend life in model organisms. We are part of a large NIH sponsored program, the Caenorhabditis Intervention Testing Program (CITP) with Gordon Lithgow (Buck Institute of Aging) and Monica Driscoll (Rutgers University) investigating the influence of genetic variation within and between species on the efficacy and robustness of these compounds. This is also one of the largest studies on the reproducibility of aging results ever conducted.
Synthetic biology of aging
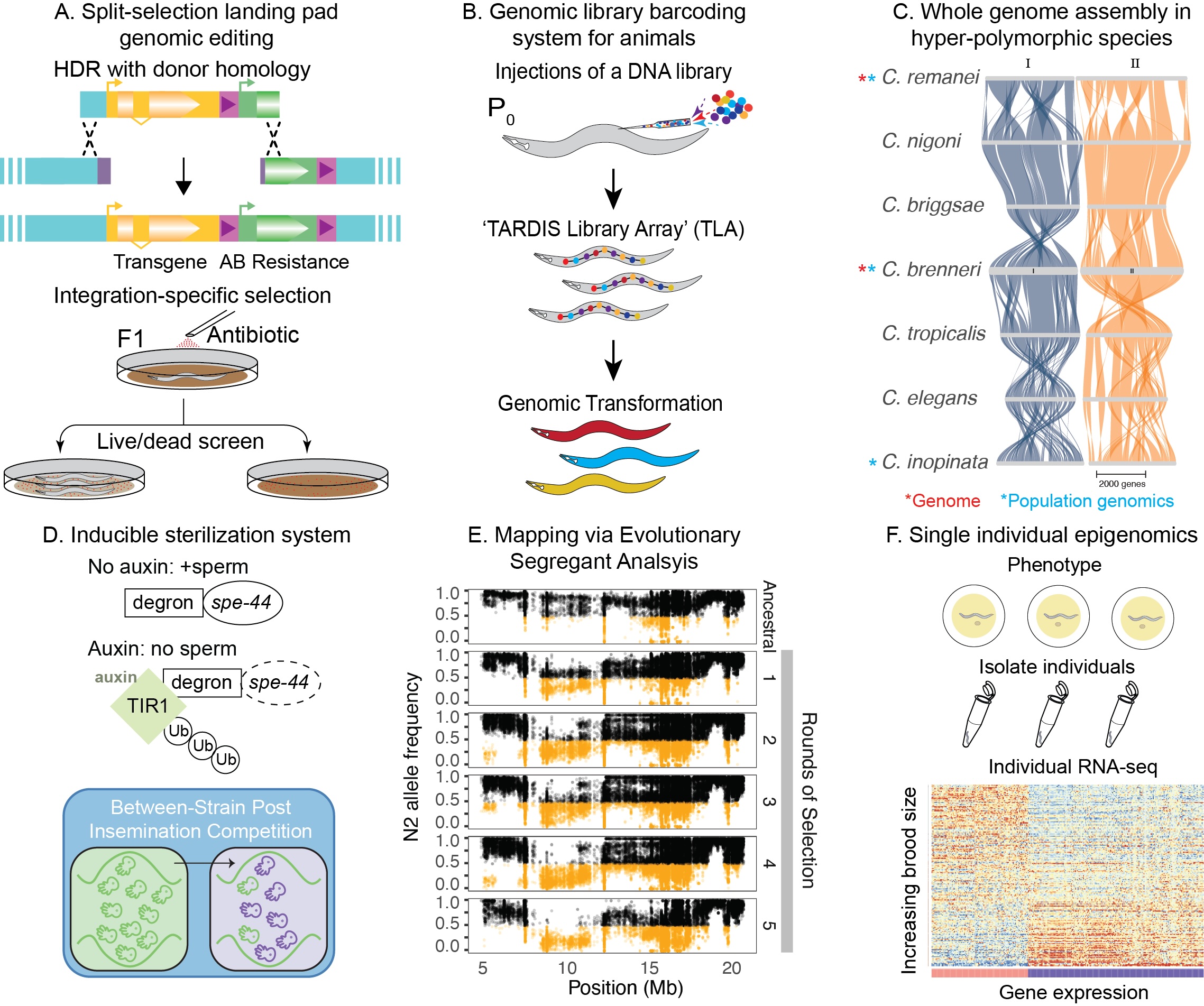
Using our recently developed library transgenesis system in C. elegans we are developing methods for directed mutagenesis and the development of synthetic genetic circuits to directly probe and manipulate genetic pathways responsible for longer life and healthy aging. In collaboration with researchers from the Knight Campus for Accelerating Scientific Impact, we are looking to extend these approaches to human cell lines and organoids, with a long term vision of greatly accelerating the discovery cycle for novel interventions that slow the rate of aging.
The genetic basis of adaptation
We have created a new genomic barcoding system that allows us to precisely trace the evolution of individual genetic lineages as they adapt to novel envrionments. This allows us to apply a variety of techniques previously only available in single-cell systems to multicellular animals. We are applying this to understand the distribution of mutational effects under differing environmental conditions but are also applying these same techniques to create new synthetic genetic systems to more explicitly understand the evolution of complex genetic networks.
Evolution of sex, outcrossing and sexual conflict
Nematodes show unusual variation in mating systems. C. elegans is androdeocious (males and hermaphrodites), while most other members of the genus are dioecious/gonochoristic (males and females). We capitalize on detailed knowledge of the C. elegans sex determination system and use experimental evolution to examine the role of males and outcrossing in facilitating evolutionary change. We also use molecular quantitative genetics to investigate what makes "good" males and females, and what allows males to be maintained at high frequencies in some populations but not in others. The high frequency of males in dioecious species like C. remanei provides a strong opportunity for sexual selection and sexual conflict, which we are investigating using genetic, genomic, proteomic, as well as more traditional behavioral, approaches.
Evolution of gene interactions and genetic networks
Standard evolutionary theory has yet to fully come to grips with the massively interactive structure of gene regulatory systems. We are looking at the evolution of genetic networks using both empirical and experimental approaches. For example, we have been looking at the molecular evolution of chemosensory, insulin signaling and stress response pathways in nematodes. We are also doing theoretical work on the evolution of network structure using whole-genome individual based simulation approaches.
Approaches
In addition to standard techniques such as genetic crosses, statistical genetics, DNA sequencing, cloning, molecular population genetics, and computer simulations, we have been active in developing novel approaches to addressing these research questions.
Experimental evolution
We have helped to pioneer the use of C. elegans and its relatives as model systems for the use of experimental evolution in asking evolutionary questions that range from the evolution of sex and outcrossing to the changes in phenotypic plasticity and transgeneration inheritence for stress response. These approaches can also be used to evolve novel biological responses using approaches from synthetic biology. Nearly every project that goes on in the lab includes some component of experimental evolution.
Population genomics
We use whole genome resequencing to analyze nematode molecular evolution. We capitalize on the work that we have done to create the first complete genome sequences of hyperpolymorphic nematode species, C. remanei and C. brenneri to push the boundaries of the current state of the art in molecular population genomics.
Functional genomics
We use a variety of protein-associated DNA approaches (e.g., ChIP-seq) and whole genome transcriptional analysis to characterize complex regulatory relationships among genes and other regulatory elements.
Synthetic biology
We have developed a number of novel large-scale genomic engineering approaches in C. elegans that allow us to systematically screen the genome for functional elements and to create novel genetic circuits to address a wide variety of biological questions, from the genetics of aging to the evolutionary consequences of sex.
Genetic mapping
We have a long history of identifying the underlying genetic basis of complex phenotypes such as behavior and stress resistance. We are applying novel whole genome bulk segregant approaches to this work, including an analysis of natural variation in extreme longevity within nematodes.
Phenomics/Microfluidics
Now that we can have total genomic knowledge regarding any individual, on of the great challenges in evolutionary biology is obtaining large amount of high quality phenotypic data on thousands of individuals. Nematodes are very small (~1 mm) and easily manipulated in liquid, making them ideal candidates for microfluidic approaches, which involve engineering liquid control circuits much like one generates computer circuits.
Parallel computing
Investigating the population genetics of large (e.g., empirically motivated) genetic networks is computationally intensive. We are developing highly parallelized approaches to addressing these questions using the new multi-million dollar computer cluster at the University of Oregon.